Proteins are the most versatile and functionally diverse macromolecules in the biological world. While DNA has the blueprint for life, proteins are the actual workers who carry out the instructions. However, a protein is not just a series of chemical components; it is a sophisticated molecular machine whose power derives entirely from its form.
The process by which a linear chain of amino acids is transformed into a complex, three-dimensional masterpiece is known as protein folding. Understanding this process is fundamental to modern biochemistry, as the rule “form follows function” dictates every breath we take, every heartbeat, and even how our bodies fight infection.
At the most basic level, proteins are polymers constructed from 20 different monomers called amino acids. Each amino acid shares a common core structure: a central carbon atom ($\alpha$-carbon) bonded to a hydrogen atom, an amino group ($-NH_2$), a carboxyl group ($-COOH$), and a unique side chain known as R group.
Synthesis of polypeptides
During the translation process in the ribosome, amino acids are linked via peptide bonds. This covalent bond is formed through a dehydration synthesis reaction between the carboxyl terminus of one amino acid and the amino terminus of the next.
The resulting chain is called a polypeptide. While the terms “polypeptide” and “protein” are often used interchangeably in casual conversation, scientists distinguish them by their condition: a polypeptide is the raw chemical chain, while a protein is a polypeptide that has folded into its functional, biologically active 3D conformation.
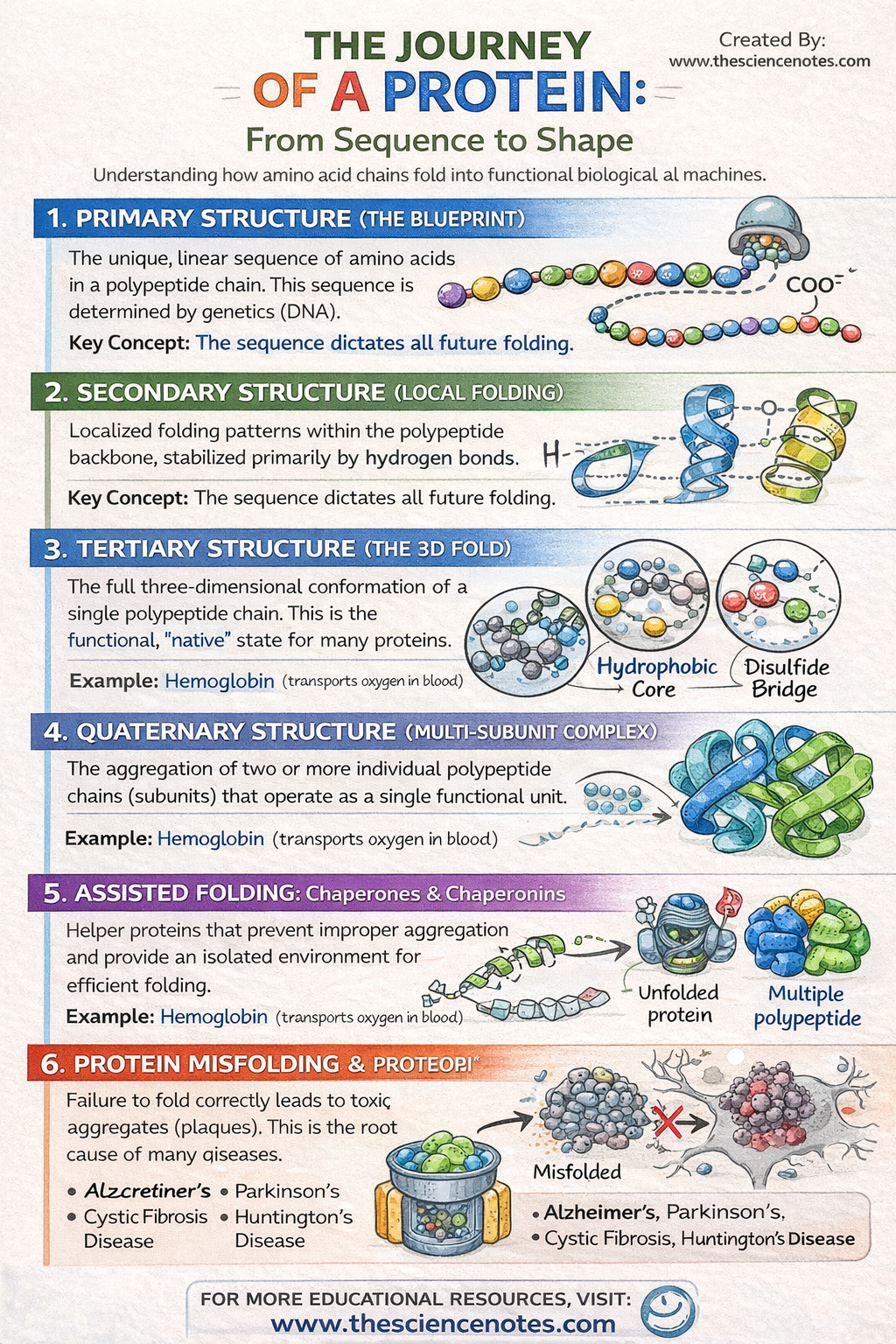
To deal with the enormous complexity of these molecules, researchers describe protein structure through four distinct hierarchical levels.
I. Primary Structure: The Genetic Blueprint
The primary structure is simply the linear sequence of amino acids. Despite its simplicity, this sequence is the most critical determinant of the protein’s future. The specific order of amino acids is dictated by the DNA sequence of the corresponding gene. Because each of the 20 amino acids has different chemical properties (size, charge, and hydrophobicity), their arrangement determines exactly how the chain will ultimately attract or repel itself to form a 3D shape.
II. Secondary structure: Localized folding
As the polypeptide exits the ribosome, it begins to form localized “neighborhoods” of shape. These are stabilized by hydrogen bonds between the atoms in the polypeptide backbone (not the side chains).
-
Alpha Helices ($\alpha$-helices): A delicate, helix-like helix held together by hydrogen bonds between every four amino acids.
-
Beta pleated sheets ($\beta$-sheet): Two or more segments of the chain lie side by side, connected by hydrogen bonds to form a rigid, sheet-like structure.
III. Tertiary Structure: The Global 3D Fold
This level represents the final “native conformation” for most single-chain proteins. While the secondary structure is about the backbone, the tertiary structure is about R group interactions. This is where the protein collapses into a globular or fibrous shape based on the chemistry of the side chains.
IV. Quaternary Structure: Multi-Unit Assemblies
Some of the most complex proteins, such as hemoglobin or DNA polymerase, consist of several polypeptide chains (subunits) that must come together to function. This assemblage is the quaternary structure. Without the proper arrangement of these subunits, the protein remains inactive.
3. The forces that drive folding
Protein folding is a “search” for the most thermodynamically stable state. Several key chemical forces act as “engineers” of this process:
The hydrophobic effect
This is perhaps the most important force in protein folding. In the cell’s aqueous environment, non-polar (hydrophobic) amino acid side chains naturally want to avoid water. When the protein folds, these hydrophobic residues cluster together in the inner “core” of the protein, while polar and charged (hydrophilic) residues remain on the outside to interact with water.
Molecular “staples”: Disulfide bonds
Cysteine is a unique amino acid because its side chain contains a sulfur-containing thiol group ($-SH$). When two cysteines are brought close together during folding, they can form a covalent bond disulfide bridge. These act as molecular staples, locking the protein in its final, most stable form and protecting it from being easily unfolded.
Van der Waals and electrostatic forces
-
Van der Waals forces: When the hydrophobic core is tightly packed, these weak attractions between the atoms provide an additional layer of structural stability.
-
Ionic bonds (salt bridges): Positively charged side chains (such as lysine) can attract negatively charged ones (such as aspartic acid) to “squeeze” parts of the protein together.
For a long time, scientists believed that proteins folded completely by themselves (Anfinsen’s dogma). However, we now know that the cellular environment is too crowded for most proteins to fold unaided. Go molecular chaperones.
-
Chaperonins: These are barrel-shaped protein complexes that act as “safe spaces”. An unfolded polypeptide enters the barrel, a “lid” closes, and the protein is allowed to fold in isolation, away from other molecules that might cause it to clump or aggregate.
-
Heat Shock Proteins (HSPs): These proteins increase in concentration when the cell is stressed by heat. They bind to exposed hydrophobic regions of unfolding proteins to prevent them from sticking together and forming toxic “clumps”.
The Role of Molecular Chaperones: The Quality Control Team
For a long time, it was believed that proteins folded completely by themselves based solely on their sequence (Anfinsen’s dogma). However, the interior of a cell is a crowded, “salty soup” of organelles and other macromolecules. In this environment, newly synthesized polypeptides have a high risk of clumping together (aggregating) or folding into “dead-end” forms that provide no biological benefit.
To ensure survival, cells have developed a sophisticated quality control system led by molecular chaperones. These proteins do not dictate the final shape of the protein – the amino acid sequence still does – but they provide the help and environment necessary for the protein to find its “native conformation” efficiently.
1. Chaperonins: The Isolation Chambers
Chaperonins, which they well studied GroEL/GroES complex in bacteria, are barrel-shaped protein structures. They act as “safe spaces” for folding.
-
Mechanism: An unfolded or partially folded polypeptide enters the central cavity of the “barrel”.
-
Insulation: A “lid” (chaperonin cap) closes the chamber. Inside this protected microenvironment, the protein is shielded from the crowded cytoplasm.
-
Folding: The environment inside the barrel often has chemical properties that favor correct folding. Once the process is complete, the lid is opened and the functional protein is released.
2. Heat Shock Proteins (HSPs): The molecular bodyguards
Heat shock proteins, such as Hsp70is the cell’s first line of defense against misfolding, especially under environmental stress such as high fever or pH changes.
-
Mechanism: They identify and bond with the vulnerable hydrophobic areas on an unfolded polypeptide.
-
Prevention: By “masking” these sticky hydrophobic patches, HSPs prevent the polypeptide from attaching to other proteins in the cell.
-
Release: Uses energy from ATPthe HSP eventually frees the protein, giving it another chance to fold correctly.
Comparison: Chaperonins vs. Heat Shock Proteins
While both are chaperones, they operate at different stages of the protein’s life cycle.
| Feature | Heat shock proteins (eg Hsp70) | Chaperonins (e.g. GroEL/ES) |
| Physical form | Small, clamp-like proteins. | Large, barrel-shaped complexes. |
| Primary action | Binds to and stabilizes “sticky” areas. | Provides an insulated “cage” for folding. |
| Timing | Often works early, while the protein is being made. | Acts later, on partially folded intermediates. |
| Energy use | Requires ATP to bind/release the protein. | Requires ATP to close lid and cycle barrel. |
| Goal | Prevents aggregation and “misfolding” under stress. | Simplifies the last “native” 3D fold. |
Enzymatic helpers: PDI and PPI
In addition to chaperones, specific enzymes accelerate the chemical “unlocking” of a protein:
-
Protein Disulfide Isomerase (PDI): This enzyme is critical for proteins that require disulfide bonds. It helps the protein quickly “test” different bond combinations until the most stable, correct disulfide bridges are formed.
-
Peptidyl Prolyl Isomerase (PPI): This enzyme helps rotate bonds involving the amino acid Prolinewhich is often a “break” in the chain that can slow down the folding process.
Without this team of chaperones and enzymes, the “folding funnel”—the path a protein takes to find its stable form—would be too slow and error-prone, leading to the cellular “garbage” that causes neurodegenerative diseases.
5. Architectural diversity: spherical vs. fibrous
Proteins generally fall into two broad structural categories based on their tertiary or quaternary forms:
Globular proteins
These are spherical, compact and generally soluble in water. Their surfaces are covered in hydrophilic residues, making them perfect for moving through the blood or cytoplasm.
-
Examples: Hemoglobin (oxygen transport), insulin (hormone signaling), and almost all enzymes (catalysis).
Fibrous proteins
These are long, rope-like and insoluble in water. They are built for strength and durability rather than chemical reactivity.
-
Examples: Keratin (strengthen hair and skin), The collagen (gives structure to tendons and bones), and Actin/Myosin (facilitating muscle movement).
6. When folding goes wrong: Denaturation and disease
Since a protein’s function is purely dependent on its shape, it loses its shape—a process called denaturation– It is usually catastrophic.
Causes of denaturation
-
Heat: Increasing kinetic energy, the protein vibrates until weak hydrogen bonds are broken.
-
pH changes: Disrupts the ionic bonds (salt bridges) by changing the charge of the side chains.
-
Chemicals: Urea or detergents can disrupt the hydrophobic core.
Proteopathy: The diseases of misfolding
If a protein misfolds and the cell’s quality control systems (such as chaperones) fail to fix or destroy it, these proteins can aggregate into “amyloid plaques.” These plaques act as “molecular sand” in the gears of the cell, eventually leading to cell death. This is the underlying mechanism for many neurodegenerative conditions:
-
Alzheimer’s disease: Caused by the accumulation of beta-amyloid plaques.
-
Parkinson’s disease: Linked to misfolding of alpha-synuclein.
-
Cystic fibrosis: Caused by a single amino acid deletion that prevents a membrane protein from folding correctly, causing it to be destroyed by the cell before it can ever function.
Conclusion: The Precision of Biological Engineering
The journey of a protein from a simple genetic sequence to a functional 3D machine is one of the most remarkable feats of biological engineering. Each interaction—from the strength of a covalent disulfide bond to the subtle “shyness” of a hydrophobic residue—is perfectly balanced to ensure that the protein can perform its life-sustaining role. As we continue to map the ‘proteome’, our understanding of these folding pathways will unlock new treatments for disease and allow us to design synthetic proteins that can solve global challenges in medicine and industry.